Abstract
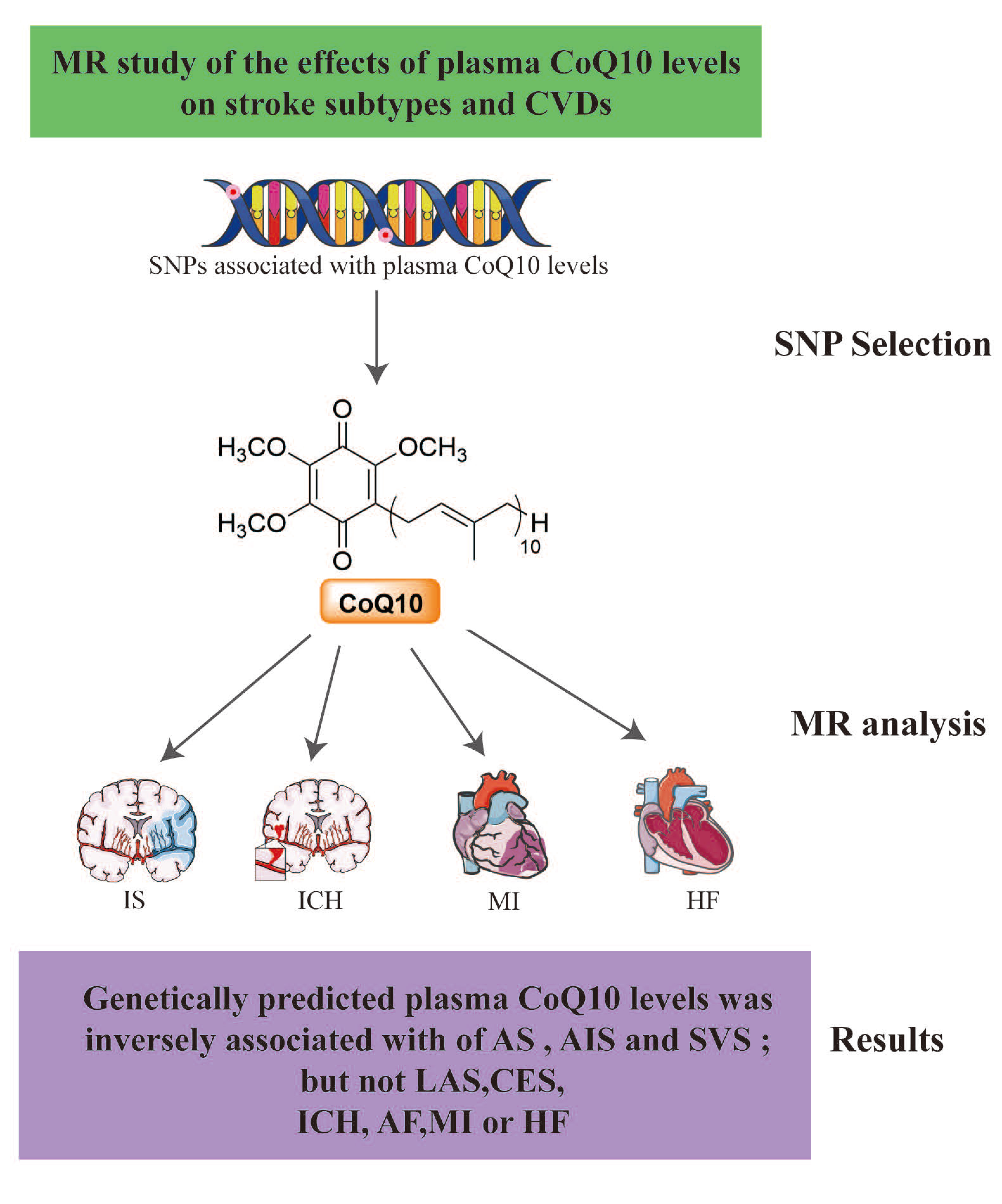
Background: Coenzyme Q10, a crucial mitochondrial compound and antioxidant, declines with age and statin use. While lower CoQ10 levels are associated with cardiovascular diseases and stroke, and supplementation shows protective effects in some heart conditions, the causal relationship between CoQ10 and stroke or cerebrovascular diseases remains unclear. The aim of this study is to investigate the causal relationship between plasma Coenzyme Q10 (CoQ10) levels and stroke, as well as some cardiovascular diseases, using Mendelian randomization methods.
Methods:Causal links were investigated through two-sample Mendelian randomization (MR) analysis. We identified genetic variants that showed significant associations with plasma CoQ10 levels. The inverse variance weighted (IVW) method was employed to estimate the effects. Additionally, sensitivity analysis was used to assess heterogeneity or pleiotropy.
Results:Our MR analysis revealed that genetically predicted plasma CoQ10 levels was inversely associated with of any stroke (AS, OR = 0.803, 95% CI: 0.659–0.978, p=0.029), any ischemic stroke (AIS, OR = 0.792, 95% CI 0.651–0.964, p = 0.020) and small vessel stroke (SVS, OR = 0.512, 95% CI 0.294-0.892, p = 0.018). However, no associations were observed between genetically predicted plasma CoQ10 and large artery stroke (LAS), cardioembolic stroke (CES), intracranial hemorrhage (ICH), atrial fibrillation (AF), myocardial infarction (MI) or heart failure (HF).
Conclusions:Our MR analysis implies a protective effect between higher plasma CoQ10 levels and AS, AIS or SVS. The results and the underlying pathways or mechanisms between plasma CoQ10 levels and stroke needs further investigation.
Keywords:
Introduction
CoQ10, also known as ubiquinone, is a naturally occurring fat-soluble compound with vitamin-like properties found in mitochondria, cell membranes, and blood [1] . Within mitochondria, CoQ10 plays a crucial role in the electron transport chain, facilitating the transfer of electrons from complexes I and II to complex III [2] . Outside mitochondria, CoQ10 is primarily distributed across various membrane structures and possesses antioxidant properties that mitigate oxidative stress damage [3,4] . However, with the aging process, the levels of coenzyme Q10 in both plasma and tissues decline significantly [5,6] . At the same time, the intake of certain medications, such as statins, can also impact the synthesis of CoQ10, leading to a reduction in its content [7] . Previous studies have indicated that supplementing CoQ10 has a certain protective effect against ischemic heart disease [8,9] , hypertension [10,11] , and heart failure [12,13] . Some studies have found a decrease in plasma CoQ10 levels in stroke patients [14] , and this decrease is correlated with the deterioration of neurological function induced by stroke [15] . Nevertheless, there is still a lack of reliable randomized controlled trial (RCT) research on the relationship between CoQ10 supplement and cerebrovascular diseases such as stroke. In addition, it is unclear whether lower plasma CoQ10 levels is a potential cause or an accompanying phenomenon of stroke and main cerebrovascular diseases.
Mendelian randomization analysis stands as a genetic epidemiological method for inferring causality. It utilizes genetic variants derived from summary-level genome-wide association study (GWAS) data as instrumental variables (IVs), effectively minimizing sampling errors and confounding factors [16] . The objective of this MR study is to assess the causal relationships between genetically predicted plasma CoQ10 levels and the susceptibility to stroke and main cardiovascular diseases.
Method
MR assumptions
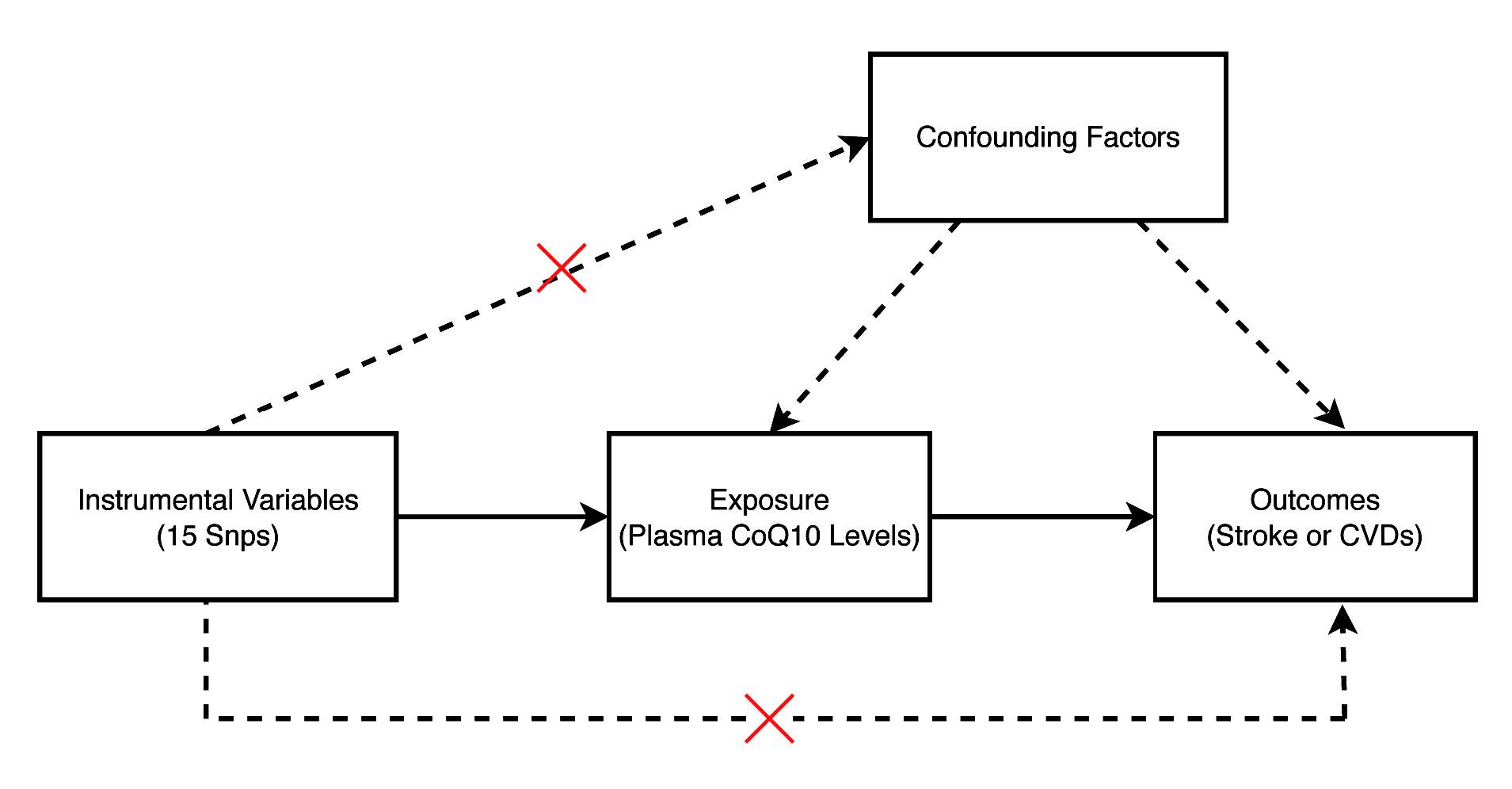
The current MR analysis relies on three primary assumptions. Firstly, the IVs are associated with the plasma CoQ10 levels at genome-wide significance; secondly, the IVs show no correlation with any potential confounding factors; thirdly, the IVs have no direct impact on stroke or cardiovascular diseases (CVDs), potentially influencing them only indirectly through plasma CoQ10 levels. The design of the present MR study is visually presented in Figure 1.
Figure 1 The main assumptions of the present Mendelian randomization study of plasma CoQ10 levels and stroke or CVDs. A total of 15 SNPs were selected as instrumental variables. CVDs, cardiovascular diseases; SNP, single nucleotide polymorphism
Plasma CoQ10 levels GWAS summary dataset.
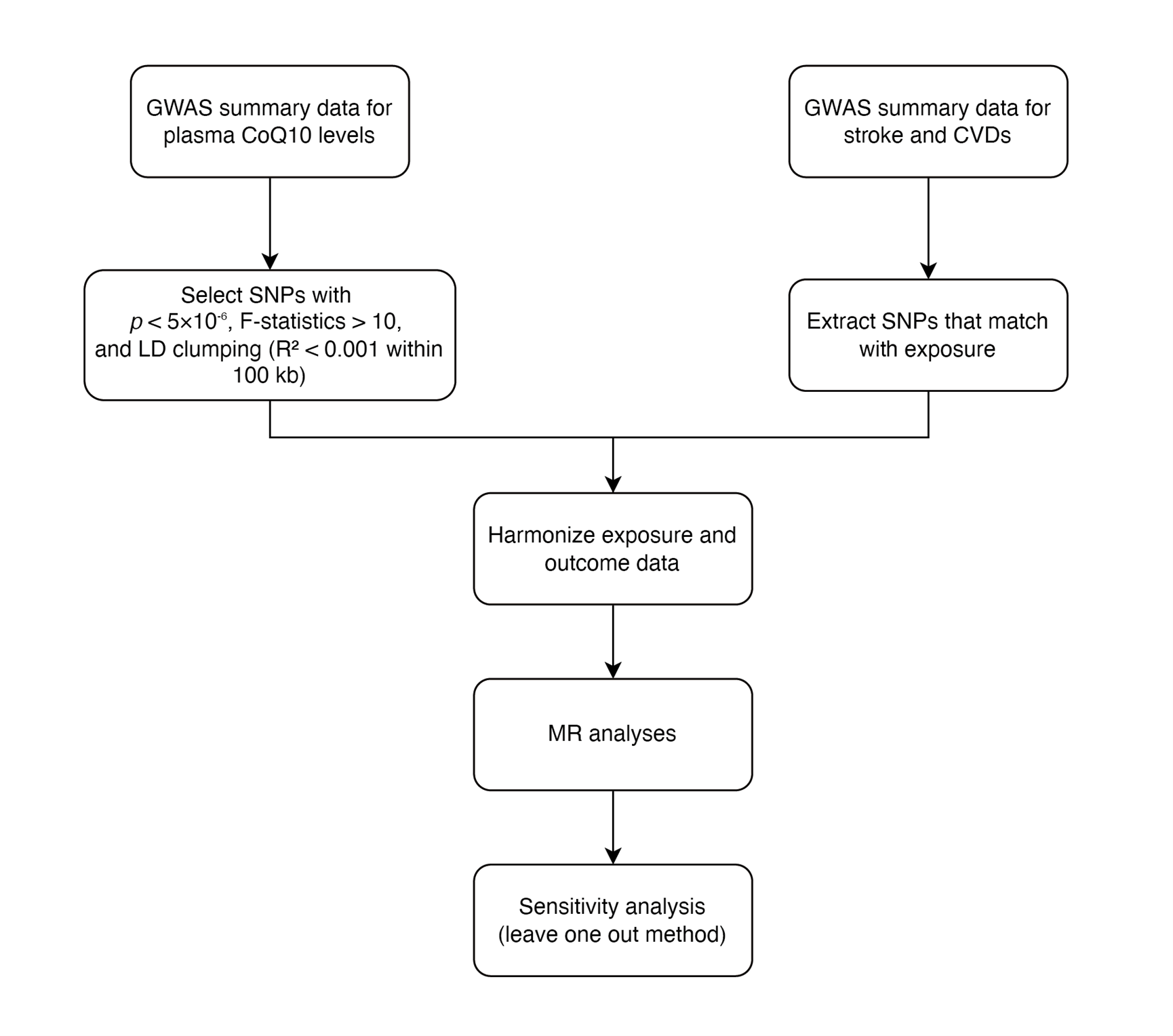
Our main analysis workflow is presented in Figure 2.We obtained genetic variants linked to plasma CoQ10 levels from a GWAS summary dataset of a 1300 cross-sectional German cohort [17] , which includes 846 participants from the PopGen cohort and 454 participants from the FoCus cohort, all European ancestry [18] . This GWAS meta-analysis was adjusted for age, sex and study center. Initially, we selected 18 SNPs that met the criterion of p < 5e-6. Subsequently, SNPs with an R2 > 0.001 within a 10,000 kb window were excluded after conducting a linkage disequilibrium test [19] . We calculated the F statistic for each SNP following the previously described method [20] . Then, SNPs with an F statistic less than 10, indicating a weak instrumental variable, were also excluded. As a result, we identified 15 SNPs that emerged as predictors for plasma CoQ10 levels (Table 1 ).
Table 1 Genetic variants significantly associated with the plasma CoQ10 levels.
|
ID |
Chr |
OA |
EA |
EAF |
β |
SE |
p |
F statistic |
|
rs9952641 |
18 |
G |
A |
0.10 |
0.063 |
0.011 |
1.31 x 10-8 |
38.40 |
|
rs933585 |
2 |
G |
A |
0.43 |
-0.034 |
0.006 |
3.64 x 10-8 |
105.08 |
|
rs686030 |
9 |
C |
A |
0.87 |
-0.044 |
0.009 |
1.39 x 10-6 |
40.58 |
|
rs35996509 |
11 |
C |
T |
0.23 |
0.037 |
0.008 |
1.92 x 10-6 |
63.82 |
|
rs150057671 |
6 |
G |
A |
0.32 |
0.034 |
0.007 |
2.44 x 10-6 |
78.72 |
|
rs12480807 |
20 |
A |
G |
0.27 |
-0.033 |
0.007 |
2.66 x 10-6 |
70.82 |
|
rs41313321 |
9 |
G |
A |
0.14 |
0.045 |
0.010 |
2.70 x 10-6 |
42.05 |
|
rs17769758 |
12 |
G |
A |
0.11 |
-0.049 |
0.010 |
3.61 x 10-6 |
33.03 |
|
rs9426691 |
1 |
A |
G |
0.27 |
-0.032 |
0.007 |
3.85 x 10-6 |
69.28 |
|
rs61745943 |
12 |
A |
T |
0.04 |
0.071 |
0.015 |
3.93 x 10-6 |
11.89 |
|
rs11591201 |
1 |
G |
A |
0.27 |
-0.034 |
0.007 |
4.02 x 10-6 |
69.30 |
|
rs7141874 |
14 |
G |
A |
0.92 |
-0.057 |
0.012 |
4.02 x 10-6 |
24.46 |
|
rs1462324 |
3 |
G |
A |
0.93 |
0.052 |
0.011 |
4.24 x 10-6 |
21.80 |
|
rs283228 |
6 |
A |
C |
0.31 |
-0.030 |
0.007 |
4.77 x 10-6 |
74.65 |
|
rs898838 |
2 |
T |
C |
0.47 |
0.028 |
0.006 |
4.84 x 10-6 |
87.48 |
Abbreviations: Chr, chromosome; EA, effect allele; EAF, effect allele frequency; OA, other allele; SE, standard error; SNP, single nucleotide polymorphism.
Figure 2 The study design of the present Mendelian randomization study of plasma CoQ10 levels and stroke or CVDs. A total of 15 SNPs were selected as instrumental variables. CVDs, cardiovascular diseases; SNP, single nucleotide polymorphism
Stroke and CVDs GWAS datasets
We acquired genetic variants for the outcomes from comprehensive GWAS datasets that encompassed main stroke subtypes and CVDs. Specifically, from the MEGASTROKE consortium, we obtained stroke-associated GWAS datasets [21] . The MEGASTROKE study included 40,585 stroke cases and 406,111 controls, categorized as follows: 34,217 cases of ischemic stroke, 4,373 cases of large vessel stroke, 7,193 cases of cardioembolic stroke, and 5,386 cases of small vessel stroke. Additionally, we obtained GWAS summary datasets for ICH, MI, HF, and AF from the FinnGen group [22] , CARDIoGRAM consortium [23] , HERMES consortium [23] , and Nielsen et al. [24] , respectively. Supplementary Table 1 provides detailed information about these datasets. All individuals included in these datasets for our MR study were of European ancestry.
Statistical analysis
We employed the IVW method as the primary tool for causal effect analysis and utilized other analytical methods [25] , such as the weighted median, weighted mode, simple median, and simple mode, as supplementary analyses. We conducted heterogeneity and pleiotropy analyses using MR-Egger, MR-PRESSO, Cochrane’s Q test, and leave-one-out analysis [26,27] . All the analyses were performed using the TwoSampleMR package.
Results
Genetically predicted plasma CoQ10 levels and risk of stroke
The IVW method revealed that genetically predicted plasma CoQ10 levels was linked to a reduced risk of stroke (OR = 0.803, 95% CI: 0.659–0.978, p = 0.029) (Figure 3 and Supplementary Figure 1,2). Subsequent analysis categorized stroke into hemorrhagic or ischemic types, we found genetically predicted plasma CoQ10 levels were inversely associated with AIS (OR = 0.792, 95% CI 0.651–0.964, p = 0.020) but not ICH (OR = 1.002, 95% CI 0.722–1.389, p = 0.992) (Figure 3 and Supplementary Figure 1,2). Further exploring the etiology of ischemic stroke indicated that genetically predicted plasma CoQ10 levels were linked to a reduced risk of SVS (OR = 0.512, 95% CI 0.294-0.892, p = 0.018) (Figure 3 and Supplementary Figure 1,2). However, no causal links were observed between genetically predicted plasma CoQ10 levels and the risk of LAS (OR = 0.917, 95% CI 0.551– 1.525, p = 0.737) or CES (OR = 1.043, 95% CI 0.715–1.523, p = 0.826) (Figure 3 and Supplementary Figure 1,2). Further details regarding the estimated effects of each SNP on stroke and its subtypes can be found in Supplementary Table 2.
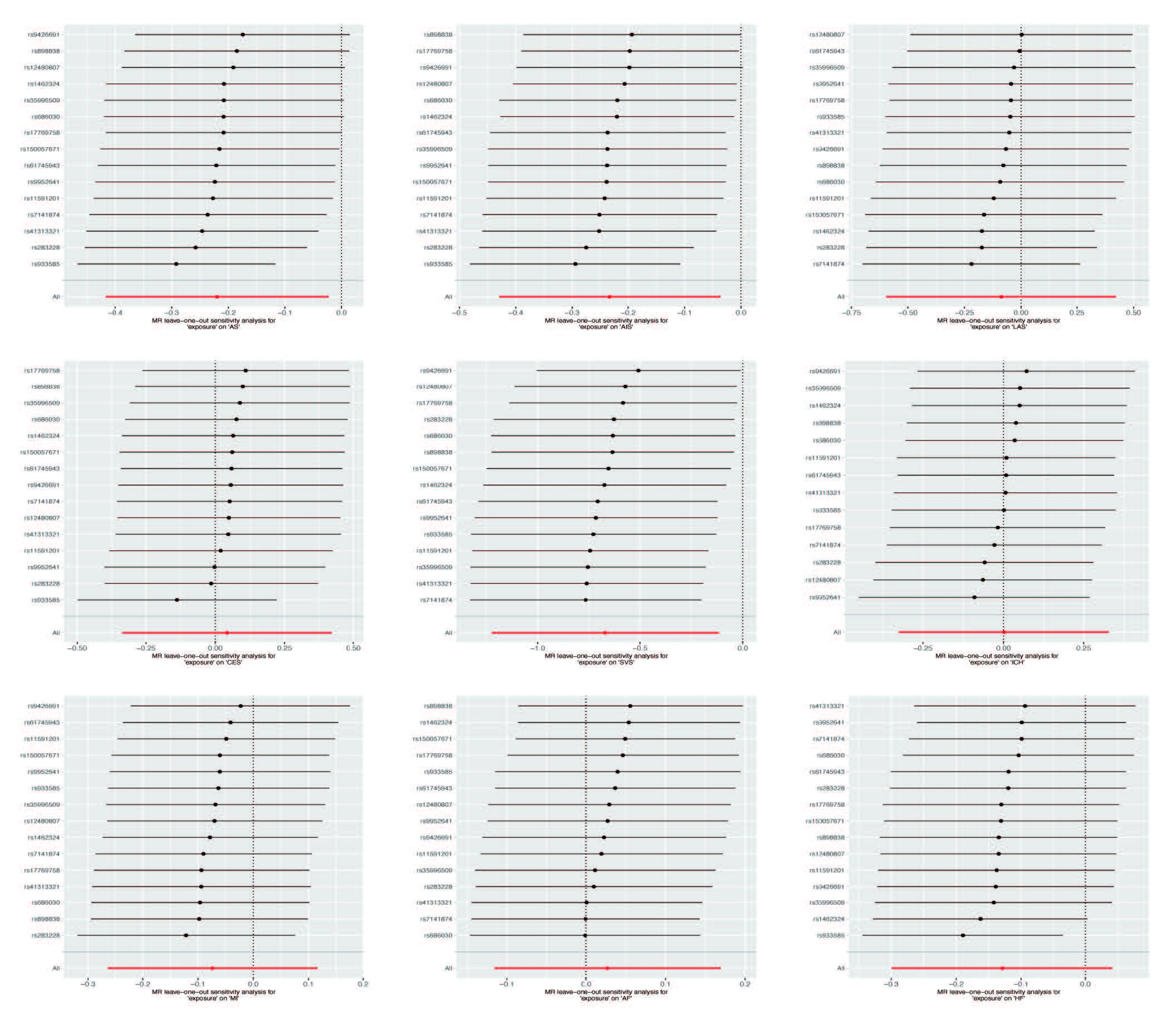
Our sensitivity analysis revealed no evidence of heterogeneities (all pval > 0.05) or pleiotropies (all pval > 0.05) in the estimated effects of plasma CoQ10 levels on stroke or its subtypes. (Supplementary Table 4). Furthermore, the leave-one-out analysis demonstrated the robustness of the MR estimated effects, as no single SNP had a substantial impact on the overall results (Figure 4).
Figure 3 Two-sample mendelian randomization analysis between plasma CoQ10 levels and stroke or cardiovascular diseases by IVW method. AF: atrial fibrillation; AS: any stroke; CES: cardio-embolic stroke; CI: confidence interval; HF: heart failure; ICH: intracranial hemorrhage; IS: ischemic stroke; IVW: inverse variance weighted; LAS: large artery atherosclerosis stroke; MI: myocardial infarction; OR: odds ratio; SNP: single nucleotide polymorphism; SVS: small vessel occlusion stroke;
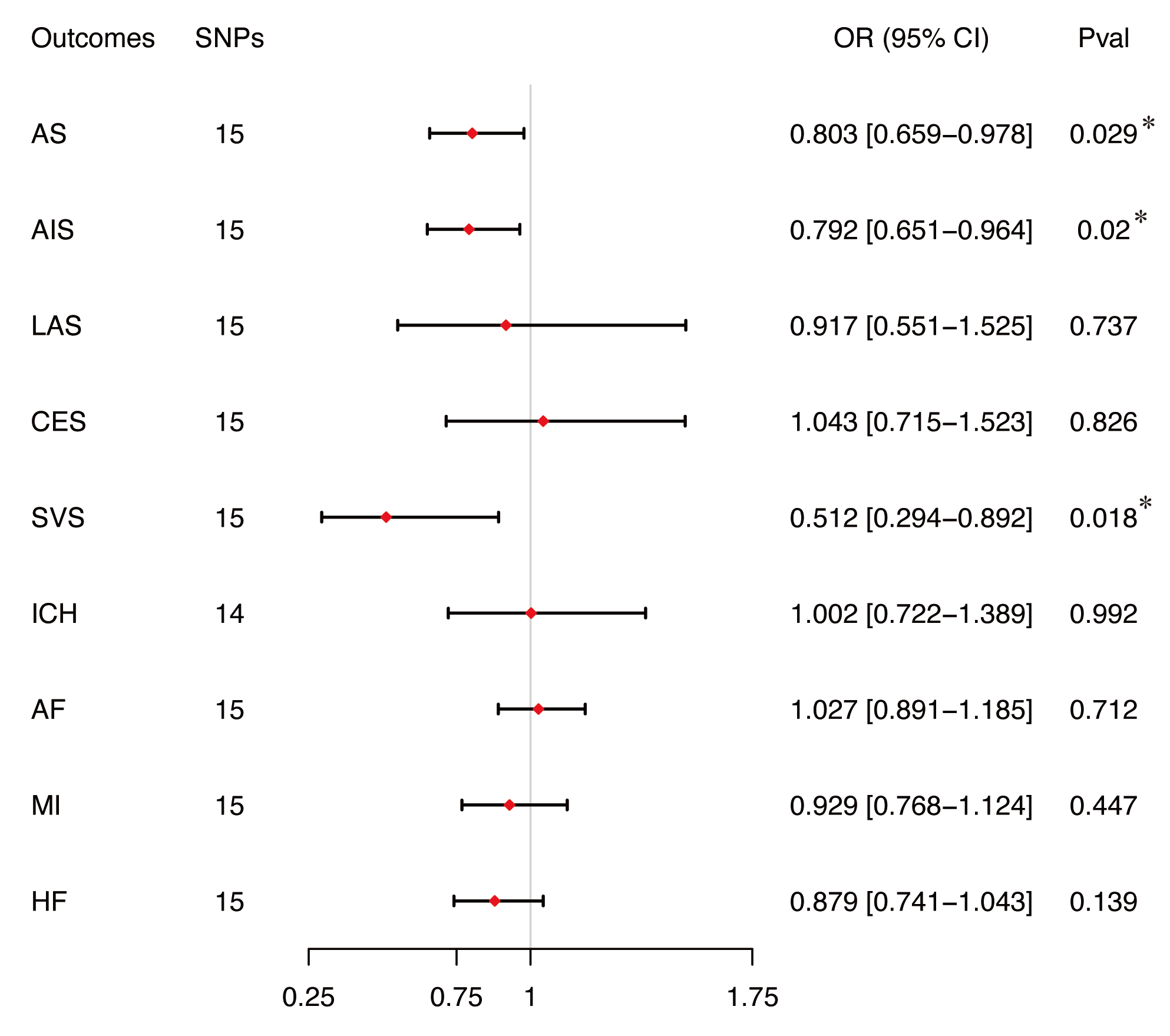
Figure 4 Sensitivity analysis of the association between genetically predicted plsma CoQ10 levels and risk of (A) AS, (B) AIS, (C) SVS, (D) LAS, (E) CES, (F) MI, (G) HF, (H) ICH, (I) AF by leave-one-out method.
In summary, our MR analysis indicates that genetically predicted plasma CoQ10 levels are inversely associated with the risk of AS, showing a 20% decrease in AS risk for each one standard deviation increase in plasma CoQ10 levels. Additionally, we observed a potential link between genetically predicted plasma CoQ10 levels and AIS or SVS, with a one standard deviation increase in quantile-normalized plasma CoQ10 levels corresponding to a 21% decreased risk of AIS and a 49% decreased risk of SVS in stroke subtypes.
Genetically predicted plasma CoQ10 levels and risk of CVDs
In general, our analysis revealed suggestive (but not statistically significant) evidence of an inverse association between genetically predicted plasma CoQ10 levels and HF risk (OR=0.879, 95% CI 0.741–1.043, p = 0.139). No significant causal link was found between genetically predicted plasma CoQ10 levels and the risk of AF (OR = 1.027, 95% CI 0.891–1.895, p = 0.712) or MI (OR = 0.929, 95% CI 0.768–1.124, p = 0.447) (Figure 3). Supplementary Table 3 provides more details regarding the estimated effects of each SNP on CVDs. Sensitivity analysis demonstrated no heterogeneities (all p > 0.05) or pleiotropies (all p > 0.05) in the estimated effects of plasma CoQ10 levels on CVDs (Supplementary Table 4). Additionally, the leave-one-out analysis indicated that the associations between plasma CoQ10 levels and CVDs were not primarily driven by any individual SNP (Figure 4).
Discussion
Coenzyme Q10, as a dietary supplement, has been widely used in the prevention and treatment of various CVDs. This study represents the first systematic exploration of the causal relationship between plasma CoQ10 levels and stroke or CVDs.
In this study, we utilized genetic variations associated with plasma CoQ10 levels as instrumental variables and employed MR to systematically analyze the causal relationship between plasma CoQ10 levels and stroke, as well as other CVDs. Our results show that (1) genetically predicted plasma CoQ10 levels were inversely associated with risk of AS and AIS; (2) regarding the etiology of ischemic stroke, genetically predicted plasma CoQ10 levels were associated with a lower risk of SAS, but not CES or LAS; (3) no causal links were identified between genetically predicted plasma CoQ10 levels and the risk of ICH, AF, MI and HF.
Our MR analysis revealed a significant protective association between genetically predicted plasma CoQ10 levels and SVS (OR=0.512, 95% CI=0.294–0.892, P=0.018), but not with CES. The relatively wide confidence interval may reflect:(1) Limited sample size of the SVS subgroup (n=6,399 cases) reducing estimate precision;(2) Effect heterogeneity in SVS pathophysiology (e.g., diverse small vessel pathologies such as arteriolosclerosis or CAA) potentially diluting IV strength;(3) Influence of outliers in the genetic instrument-exposure association. Despite the wide CI, the significant protective effect (OR<1, P<0.05) is consistent across sensitivity analyses, supporting further validation in larger cohorts.
Notwithstanding these statistical considerations, the observed subtype-specific protective effect for SVS warrants mechanistic interpretation. This specificity may stem from distinct pathophysiological mechanisms across stroke subtypes. SVS is primarily characterized by cerebral small vessel endothelial dysfunction and blood-brain barrier disruption, where oxidative stress plays a central role. CoQ10, as a potent mitochondrial electron transporter and lipid-soluble antioxidant, may mitigate oxidative damage in small vessel endothelial cells more effectively than in larger vessels. Small vessels exhibit higher mitochondrial density and metabolic activity, rendering their endothelium particularly vulnerable to reactive oxygen species (ROS)-induced injury. The observed protective effect could thus reflect CoQ10's ability to stabilize endothelial function and reduce oxidative stress in microvasculature, aligning with its known role in suppressing ROS generation and preserving nitric oxide (NO) bioavailability.
In contrast, LAS and CES involve thromboembolic mechanisms (e.g., atherosclerotic plaque rupture or atrial fibrillation-related clots) where oxidative stress may be less dominant. Additionally, the null association with intracranial hemorrhage (ICH) (OR=1.002, p=0.992) suggests CoQ10's effects are specific to ischemic pathways rather than hemorrhagic pathophysiology.
The absence of causal links between CoQ10 and cardiovascular outcomes like AF, MI, HF may reflect disease-specific mechanisms. For AF, electrical remodeling and fibrosis are driven more by ion channel dysfunction and inflammation than mitochondrial oxidative stress. Similarly, MI pathogenesis centers on acute plaque rupture and thrombosis, where CoQ10's antioxidant properties may not sufficiently counteract abrupt thrombotic events. Although prior RCTs suggest CoQ10 supplementation benefits HF patients, our null genetic association (OR=0.879, p=0.139) implies that plasma CoQ10 levels alone may not modulate HF risk through the pathways captured by our instrumental variables. This discrepancy could arise from differences in study design (e.g., MR assesses lifelong exposure vs. RCTs examining short-term supplementation).
Previous studies have reported associations between CoQ10 and stroke. It was found that stroke patients have lower plasma CoQ10 levels compared to the normal control group [14] . Additionally, early supplementation of CoQ10 in patients after acute ischemic stroke can improve neurological function scores, as measured by the National Institute of Health Stroke Scale (NIHSS) [15] . Our research further indicates a potential association between the plasma CoQ10 levels and the occurrence of stroke. This effect may be related to the significant antioxidant effect of CoQ10, which protects the vascular endothelium, prevents dysfunction of endothelial cells, and helps in the prevention of stroke [28,29] . Due to the gradual decline in CoQ10 levels with increasing age, this decrease may become an important contributing factor to the occurrence of strokes in the elderly population. Previous studies have already demonstrated the supplemental role of CoQ10 in the adjunctive treatment of cardiovascular diseases such as hypertension and heart failure. This study further suggests that moderate supplementation of CoQ10 in the elderly population may provide a certain protective effect against the occurrence of strokes.
This study also has certain limitations. Due to the lack of large-scale GWAS results on plasma CoQ10, there may be a presence of weak instrument bias. However, F statistics for these exposures were well over 10, meaning that the magnitude of bias is likely to be small. Lastly, only people of European ancestry were included in the MR analysis, which limited the generalizability of the findings of the present MR study. The proportions of stroke etiologies vary based on race or ethnicity.
Conclusions
We found genetically predicted plasma CoQ10 levels to be inversely associated with AS, AIS and SVS. Our analysis did not identify any causal associations between genetically predicted plasma CoQ10 levels and CES, LAS, AF, MI and HF.
Supplementary Material
Not applicable.
Declarations
Author contributions
Xuyang Liu conceived the analysis and drafted this manuscript. Yuyuan Gao and Xu Li were responsible for data collection. Xiaojing Yao executed data analysis. Yuming Xu contributed to the study concept and supervision. All authors have given their approval for the submitted version.
Acknowledgements
The author thanks the MEGASTROKE consortium,FinnGen, CARDIoGRAMplusC4D Consortium, Heart Failure Molecular Epidemiology for Therapeutic Targets consortium (HERMES), Nielsen et al. for making their data sets publicly available.
Funding information
This study was funded by the NHC Key Laboratory of Prevention and Treatment of Cerebrovascular Disease, Henan Key Laboratory of Cerebrovascular Diseases (Zhengzhou University), the Non-profit Central Research Institute Fund of Chinese Academy of Medical Sciences (Grant No. 2020-PT310-01), and Technology Projects of Henan Province in 2020 (Grant No. 201300310300).
Ethics approval and consent to participate
The present study was based on the secondary analysis of data publicly available and ethics consent was not required.
Competing Interests
The authors declare that they have no existing or potential commercial or financial relationships that could create a conflict of interest at the time of conducting this study.
Data Availability
All data needed to evaluate the conclusions in the paper are present in the paper or the Supplementary Materials. Additional data related to this paper may be requested from the authors.
References
[1] Testai L, Martelli A, Flori L, Cicero AFG, & Colletti A. (2021). Coenzyme Q(10): Clinical Applications beyond Cardiovascular Diseases. Nutrients, 13(5). https://doi.org/10.3390/nu13051697
[2] Ernster L, & Dallner G. (1995). Biochemical, physiological and medical aspects of ubiquinone function. Biochim Biophys Acta, 1271(1), 195-204. https://doi.org/10.1016/0925-4439(95)00028-3
[3] Bullón P, Román-Malo L, Marín-Aguilar F, Alvarez-Suarez JM, Giampieri F, Battino M, et al. (2015). Lipophilic antioxidants prevent lipopolysaccharide-induced mitochondrial dysfunction through mitochondrial biogenesis improvement. Pharmacol Res, 91, 1-8. https://doi.org/10.1016/j.phrs.2014.10.007
[4] Kernt M, Hirneiss C, Neubauer AS, Ulbig MW, & Kampik A. (2010). Coenzyme Q10 prevents human lens epithelial cells from light-induced apoptotic cell death by reducing oxidative stress and stabilizing BAX / Bcl-2 ratio. Acta Ophthalmol, 88(3), e78-86. https://doi.org/10.1111/j.1755-3768.2010.01875.x
[5] Navas P, Cascajo MV, Alcázar-Fabra M, Hernández-Camacho JD, Sánchez-Cuesta A, Rodríguez ABC, et al. (2021). Secondary CoQ(10) deficiency, bioenergetics unbalance in disease and aging. Biofactors, 47(4), 551-569. https://doi.org/10.1002/biof.1733
[6] Gasmi A, Bjørklund G, Mujawdiya PK, Semenova Y, Piscopo S, & Peana M. (2024). Coenzyme Q(10) in aging and disease. Crit Rev Food Sci Nutr, 64(12), 3907-3919. https://doi.org/10.1080/10408398.2022.2137724
[7] Langsjoen PH, & Langsjoen AM. (2003). The clinical use of HMG CoA-reductase inhibitors and the associated depletion of coenzyme Q10. A review of animal and human publications. Biofactors, 18(1-4), 101-111. https://doi.org/10.1002/biof.5520180212
[8] Ayer A, Macdonald P, & Stocker R. (2015). CoQ₁₀ Function and Role in Heart Failure and Ischemic Heart Disease. Annu Rev Nutr, 35, 175-213. https://doi.org/10.1146/annurev-nutr-071714-034258
[9] Ayers J, Cook J, Koenig RA, Sisson EM, & Dixon DL. (2018). Recent Developments in the Role of Coenzyme Q10 for Coronary Heart Disease: a Systematic Review. Curr Atheroscler Rep, 20(6), 29. https://doi.org/10.1007/s11883-018-0730-1
[10] Digiesi V, Cantini F, Oradei A, Bisi G, Guarino GC, Brocchi A, et al. (1994). Coenzyme Q10 in essential hypertension. Mol Aspects Med, 15 Suppl, s257-263. https://doi.org/10.1016/0098-2997(94)90036-1
[11] Gao HL, Yu XJ, Qi J, Yi QY, Jing WH, Sun WY, et al. (2016). Oral CoQ10 attenuates high salt-induced hypertension by restoring neurotransmitters and cytokines in the hypothalamic paraventricular nucleus. Sci Rep, 6, 30301. https://doi.org/10.1038/srep30301
[12] Mortensen SA, Rosenfeldt F, Kumar A, Dolliner P, Filipiak KJ, Pella D, et al. (2014). The effect of coenzyme Q10 on morbidity and mortality in chronic heart failure: results from Q-SYMBIO: a randomized double-blind trial. JACC Heart Fail, 2(6), 641-649. https://doi.org/10.1016/j.jchf.2014.06.008
[13] Baggio E, Gandini R, Plancher AC, Passeri M, & Carmosino G. (1994). Italian multicenter study on the safety and efficacy of coenzyme Q10 as adjunctive therapy in heart failure. CoQ10 Drug Surveillance Investigators. Mol Aspects Med, 15 Suppl, s287-294. https://doi.org/10.1016/0098-2997(94)90040-x
[14] Simani L, Ryan F, Hashemifard S, Hooshmandi E, Madahi M, Sahraei Z, et al. (2018). Serum Coenzyme Q10 Is Associated with Clinical Neurological Outcomes in Acute Stroke Patients. J Mol Neurosci, 66(1), 53-58. https://doi.org/10.1007/s12031-018-1115-1
[15] Ramezani M, Sahraei Z, Simani L, Heydari K, & Shahidi F. (2020). Coenzyme Q10 supplementation in acute ischemic stroke: Is it beneficial in short-term administration? Nutr Neurosci, 23(8), 640-645. https://doi.org/10.1080/1028415x.2018.1541269
[16] Lawlor DA, Harbord RM, Sterne JA, Timpson N, & Davey Smith G. (2008). Mendelian randomization: using genes as instruments for making causal inferences in epidemiology. Stat Med, 27(8), 1133-1163. https://doi.org/10.1002/sim.3034
[17] Degenhardt F, Niklowitz P, Szymczak S, Jacobs G, Lieb W, Menke T, et al. (2016). Genome-wide association study of serum coenzyme Q10 levels identifies susceptibility loci linked to neuronal diseases. Hum Mol Genet, 25(13), 2881-2891. https://doi.org/10.1093/hmg/ddw134
[18] Onur S, Niklowitz P, Fischer A, Jacobs G, Lieb W, Laudes M, et al. (2015). Determination of the coenzyme Q10 status in a large Caucasian study population. Biofactors, 41(4), 211-221. https://doi.org/10.1002/biof.1216
[19] Burgess S, & Thompson SG. (2011). Avoiding bias from weak instruments in Mendelian randomization studies. Int J Epidemiol, 40(3), 755-764. https://doi.org/10.1093/ije/dyr036
[20] Hemani G, Zheng J, Elsworth B, Wade KH, Haberland V, Baird D, et al. (2018). The MR-Base platform supports systematic causal inference across the human phenome. Elife, 7. https://doi.org/10.7554/eLife.34408
[21] Malik R, Chauhan G, Traylor M, Sargurupremraj M, Okada Y, Mishra A, et al. (2018). Multiancestry genome-wide association study of 520,000 subjects identifies 32 loci associated with stroke and stroke subtypes. Nat Genet, 50(4), 524-537. https://doi.org/10.1038/s41588-018-0058-3
[22] Kurki MI, Karjalainen J, Palta P, Sipilä TP, Kristiansson K, Donner KM, et al. (2023). FinnGen provides genetic insights from a well-phenotyped isolated population. Nature, 613(7944), 508-518. https://doi.org/10.1038/s41586-022-05473-8
[23] Nikpay M, Goel A, Won HH, Hall LM, Willenborg C, Kanoni S, et al. (2015). A comprehensive 1,000 Genomes-based genome-wide association meta-analysis of coronary artery disease. Nat Genet, 47(10), 1121-1130. https://doi.org/10.1038/ng.3396
[24] Nielsen JB, Thorolfsdottir RB, Fritsche LG, Zhou W, Skov MW, Graham SE, et al. (2018). Biobank-driven genomic discovery yields new insight into atrial fibrillation biology. Nat Genet, 50(9), 1234-1239. https://doi.org/10.1038/s41588-018-0171-3
[25] Burgess S, Small DS, & Thompson SG. (2017). A review of instrumental variable estimators for Mendelian randomization. Stat Methods Med Res, 26(5), 2333-2355. https://doi.org/10.1177/0962280215597579
[26] Bowden J, Davey Smith G, & Burgess S. (2015). Mendelian randomization with invalid instruments: effect estimation and bias detection through Egger regression. Int J Epidemiol, 44(2), 512-525. https://doi.org/10.1093/ije/dyv080
[27] Verbanck M, Chen CY, Neale B, & Do R. (2018). Detection of widespread horizontal pleiotropy in causal relationships inferred from Mendelian randomization between complex traits and diseases. Nat Genet, 50(5), 693-698. https://doi.org/10.1038/s41588-018-0099-7
[28] Hamilton SJ, Chew GT, & Watts GF. (2009). Coenzyme Q10 improves endothelial dysfunction in statin-treated type 2 diabetic patients. Diabetes Care, 32(5), 810-812. https://doi.org/10.2337/dc08-1736
[29] Tsai KL, Huang YH, Kao CL, Yang DM, Lee HC, Chou HY, et al. (2012). A novel mechanism of coenzyme Q10 protects against human endothelial cells from oxidative stress-induced injury by modulating NO-related pathways. J Nutr Biochem, 23(5), 458-468. https://doi.org/10.1016/j.jnutbio.2011.01.011
 Figures
Figures References
References Peer
Peer Information
InformationFigure 1 The main assumptions of the present Mendelian randomization study of plasma CoQ10 levels and stroke or CVDs. A total of 15 SNPs were selected as instrumental variables. CVDs, cardiovascular diseases; SNP, single nucleotide polymorphism

Figure 2 The study design of the present Mendelian randomization study of plasma CoQ10 levels and stroke or CVDs. A total of 15 SNPs were selected as instrumental variables. CVDs, cardiovascular diseases; SNP, single nucleotide polymorphism

Figure 3 Two-sample mendelian randomization analysis between plasma CoQ10 levels and stroke or cardiovascular diseases by IVW method. AF: atrial fibrillation; AS: any stroke; CES: cardio-embolic stroke; CI: confidence interval; HF: heart failure; ICH: intracranial hemorrhage; IS: ischemic stroke; IVW: inverse variance weighted; LAS: large artery atherosclerosis stroke; MI: myocardial infarction; OR: odds ratio; SNP: single nucleotide polymorphism; SVS: small vessel occlusion stroke;

Figure 4 Sensitivity analysis of the association between genetically predicted plsma CoQ10 levels and risk of (A) AS, (B) AIS, (C) SVS, (D) LAS, (E) CES, (F) MI, (G) HF, (H) ICH, (I) AF by leave-one-out method.

[1] Testai L, Martelli A, Flori L, Cicero AFG, & Colletti A. (2021). Coenzyme Q(10): Clinical Applications beyond Cardiovascular Diseases. Nutrients, 13(5). https://doi.org/10.3390/nu13051697
[2] Ernster L, & Dallner G. (1995). Biochemical, physiological and medical aspects of ubiquinone function. Biochim Biophys Acta, 1271(1), 195-204. https://doi.org/10.1016/0925-4439(95)00028-3
[3] Bullón P, Román-Malo L, Marín-Aguilar F, Alvarez-Suarez JM, Giampieri F, Battino M, et al. (2015). Lipophilic antioxidants prevent lipopolysaccharide-induced mitochondrial dysfunction through mitochondrial biogenesis improvement. Pharmacol Res, 91, 1-8. https://doi.org/10.1016/j.phrs.2014.10.007
[4] Kernt M, Hirneiss C, Neubauer AS, Ulbig MW, & Kampik A. (2010). Coenzyme Q10 prevents human lens epithelial cells from light-induced apoptotic cell death by reducing oxidative stress and stabilizing BAX / Bcl-2 ratio. Acta Ophthalmol, 88(3), e78-86. https://doi.org/10.1111/j.1755-3768.2010.01875.x
[5] Navas P, Cascajo MV, Alcázar-Fabra M, Hernández-Camacho JD, Sánchez-Cuesta A, Rodríguez ABC, et al. (2021). Secondary CoQ(10) deficiency, bioenergetics unbalance in disease and aging. Biofactors, 47(4), 551-569. https://doi.org/10.1002/biof.1733
[6] Gasmi A, Bjørklund G, Mujawdiya PK, Semenova Y, Piscopo S, & Peana M. (2024). Coenzyme Q(10) in aging and disease. Crit Rev Food Sci Nutr, 64(12), 3907-3919. https://doi.org/10.1080/10408398.2022.2137724
[7] Langsjoen PH, & Langsjoen AM. (2003). The clinical use of HMG CoA-reductase inhibitors and the associated depletion of coenzyme Q10. A review of animal and human publications. Biofactors, 18(1-4), 101-111. https://doi.org/10.1002/biof.5520180212
[8] Ayer A, Macdonald P, & Stocker R. (2015). CoQ₁₀ Function and Role in Heart Failure and Ischemic Heart Disease. Annu Rev Nutr, 35, 175-213. https://doi.org/10.1146/annurev-nutr-071714-034258
[9] Ayers J, Cook J, Koenig RA, Sisson EM, & Dixon DL. (2018). Recent Developments in the Role of Coenzyme Q10 for Coronary Heart Disease: a Systematic Review. Curr Atheroscler Rep, 20(6), 29. https://doi.org/10.1007/s11883-018-0730-1
[10] Digiesi V, Cantini F, Oradei A, Bisi G, Guarino GC, Brocchi A, et al. (1994). Coenzyme Q10 in essential hypertension. Mol Aspects Med, 15 Suppl, s257-263. https://doi.org/10.1016/0098-2997(94)90036-1
[11] Gao HL, Yu XJ, Qi J, Yi QY, Jing WH, Sun WY, et al. (2016). Oral CoQ10 attenuates high salt-induced hypertension by restoring neurotransmitters and cytokines in the hypothalamic paraventricular nucleus. Sci Rep, 6, 30301. https://doi.org/10.1038/srep30301
[12] Mortensen SA, Rosenfeldt F, Kumar A, Dolliner P, Filipiak KJ, Pella D, et al. (2014). The effect of coenzyme Q10 on morbidity and mortality in chronic heart failure: results from Q-SYMBIO: a randomized double-blind trial. JACC Heart Fail, 2(6), 641-649. https://doi.org/10.1016/j.jchf.2014.06.008
[13] Baggio E, Gandini R, Plancher AC, Passeri M, & Carmosino G. (1994). Italian multicenter study on the safety and efficacy of coenzyme Q10 as adjunctive therapy in heart failure. CoQ10 Drug Surveillance Investigators. Mol Aspects Med, 15 Suppl, s287-294. https://doi.org/10.1016/0098-2997(94)90040-x
[14] Simani L, Ryan F, Hashemifard S, Hooshmandi E, Madahi M, Sahraei Z, et al. (2018). Serum Coenzyme Q10 Is Associated with Clinical Neurological Outcomes in Acute Stroke Patients. J Mol Neurosci, 66(1), 53-58. https://doi.org/10.1007/s12031-018-1115-1
[15] Ramezani M, Sahraei Z, Simani L, Heydari K, & Shahidi F. (2020). Coenzyme Q10 supplementation in acute ischemic stroke: Is it beneficial in short-term administration? Nutr Neurosci, 23(8), 640-645. https://doi.org/10.1080/1028415x.2018.1541269
[16] Lawlor DA, Harbord RM, Sterne JA, Timpson N, & Davey Smith G. (2008). Mendelian randomization: using genes as instruments for making causal inferences in epidemiology. Stat Med, 27(8), 1133-1163. https://doi.org/10.1002/sim.3034
[17] Degenhardt F, Niklowitz P, Szymczak S, Jacobs G, Lieb W, Menke T, et al. (2016). Genome-wide association study of serum coenzyme Q10 levels identifies susceptibility loci linked to neuronal diseases. Hum Mol Genet, 25(13), 2881-2891. https://doi.org/10.1093/hmg/ddw134
[18] Onur S, Niklowitz P, Fischer A, Jacobs G, Lieb W, Laudes M, et al. (2015). Determination of the coenzyme Q10 status in a large Caucasian study population. Biofactors, 41(4), 211-221. https://doi.org/10.1002/biof.1216
[19] Burgess S, & Thompson SG. (2011). Avoiding bias from weak instruments in Mendelian randomization studies. Int J Epidemiol, 40(3), 755-764. https://doi.org/10.1093/ije/dyr036
[20] Hemani G, Zheng J, Elsworth B, Wade KH, Haberland V, Baird D, et al. (2018). The MR-Base platform supports systematic causal inference across the human phenome. Elife, 7. https://doi.org/10.7554/eLife.34408
[21] Malik R, Chauhan G, Traylor M, Sargurupremraj M, Okada Y, Mishra A, et al. (2018). Multiancestry genome-wide association study of 520,000 subjects identifies 32 loci associated with stroke and stroke subtypes. Nat Genet, 50(4), 524-537. https://doi.org/10.1038/s41588-018-0058-3
[22] Kurki MI, Karjalainen J, Palta P, Sipilä TP, Kristiansson K, Donner KM, et al. (2023). FinnGen provides genetic insights from a well-phenotyped isolated population. Nature, 613(7944), 508-518. https://doi.org/10.1038/s41586-022-05473-8
[23] Nikpay M, Goel A, Won HH, Hall LM, Willenborg C, Kanoni S, et al. (2015). A comprehensive 1,000 Genomes-based genome-wide association meta-analysis of coronary artery disease. Nat Genet, 47(10), 1121-1130. https://doi.org/10.1038/ng.3396
[24] Nielsen JB, Thorolfsdottir RB, Fritsche LG, Zhou W, Skov MW, Graham SE, et al. (2018). Biobank-driven genomic discovery yields new insight into atrial fibrillation biology. Nat Genet, 50(9), 1234-1239. https://doi.org/10.1038/s41588-018-0171-3
[25] Burgess S, Small DS, & Thompson SG. (2017). A review of instrumental variable estimators for Mendelian randomization. Stat Methods Med Res, 26(5), 2333-2355. https://doi.org/10.1177/0962280215597579
[26] Bowden J, Davey Smith G, & Burgess S. (2015). Mendelian randomization with invalid instruments: effect estimation and bias detection through Egger regression. Int J Epidemiol, 44(2), 512-525. https://doi.org/10.1093/ije/dyv080
[27] Verbanck M, Chen CY, Neale B, & Do R. (2018). Detection of widespread horizontal pleiotropy in causal relationships inferred from Mendelian randomization between complex traits and diseases. Nat Genet, 50(5), 693-698. https://doi.org/10.1038/s41588-018-0099-7
[28] Hamilton SJ, Chew GT, & Watts GF. (2009). Coenzyme Q10 improves endothelial dysfunction in statin-treated type 2 diabetic patients. Diabetes Care, 32(5), 810-812. https://doi.org/10.2337/dc08-1736
[29] Tsai KL, Huang YH, Kao CL, Yang DM, Lee HC, Chou HY, et al. (2012). A novel mechanism of coenzyme Q10 protects against human endothelial cells from oxidative stress-induced injury by modulating NO-related pathways. J Nutr Biochem, 23(5), 458-468. https://doi.org/10.1016/j.jnutbio.2011.01.011
Peer-review Terminology
Identity transparency: Single anonymized
Reviewer interacts with: Editor
Details
This is an open access article under the terms of the Creative Commons Attribution License(http://creativecommons.org/licenses/by/4.0/), which permits use, distribution and reproduction in any medium, provided the original work is properly cited.
Publication History
Received 2025-04-14
Accepted 2025-06-17
Published 2025-07-18


